Aim:
Single biomarker diagnostic test of BRAFV600 locus in metastatic melanoma is mandatory for treatment decision; however, multiple-gene based techniques, such as targeted next-generation sequencing (NGS) are being used to maximize the number of patients that can benefit from a targeted therapy. The main objective of this study is to investigate whether an NGS panel could be adopted in routine clinical care for advanced melanoma.
Methods:
Patients diagnosed with advanced melanoma at our center from 2017 to 2019 were included. Presence of genetic alterations was performed using two methods: real-time polymerase chain reaction-based Idylla test (Biocartis) and NGS with the oncomine solid tumor DNA kit (Thermo Fisher Scientific). Total genomic DNA was extracted from formalin-fixed and paraffin embedded samples for sequencing.
Results:
A total of 155 samples were evaluated for molecular analysis but 40 samples (25.8%) were inadequate for sequencing. The clinical utility of BRAFV600 real-time polymerase chain reaction and targeted-NGS was compared in 29 samples and a very good concordance was observed (Kappa = 0.89, 95% confidence interval 0.68 ± 1.05). An oncogenic mutation by NGS was found in 75 samples (65%)–53% of whom were candidates for personalized therapies. The most prevalent mutated genes were BRAF (39%), TP53 (23%), and NRAS (14%). Other genes identified at lower incidence (< 5%) were: PIK3CA, ERBB4, CTNNB1, STK11, FGFR1, SMAD4, KRAS, FGFR3, PTEN and AKT. Co-occurrence of oncogenic mutations was detected in 40% of the samples. Among the mutations identified, TP53 was significantly more prevalent in men (men 31.8% versus women 12.2%, P = 0.03) and NRAS in women (men 9.1% versus women 24.4%, P = 0.03).
Conclusions:
Targeted-NGS testing is a feasible technique to implement in the routine clinical practice. Based on our results, NGS has provided more information on target-genes than RT-PCR technique, maximizing the benefit for patients with advanced melanoma.
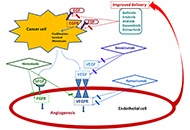 Angiogenesis and epidermal growth factor receptor inhibitors in non-small cell lung cancerOpen AccessReviewSeveral preclinical studies suggested a potential benefit from combined treatment with inhibitors of epidermal growth factor receptor (EGFR) and angiogenesis, both effective in patients with advance [...] Read more.Giuliano Palumbo ... Alessandro MorabitoPublished: April 28, 2020 Explor Target Antitumor Ther. 2020;1:117–130
Angiogenesis and epidermal growth factor receptor inhibitors in non-small cell lung cancerOpen AccessReviewSeveral preclinical studies suggested a potential benefit from combined treatment with inhibitors of epidermal growth factor receptor (EGFR) and angiogenesis, both effective in patients with advance [...] Read more.Giuliano Palumbo ... Alessandro MorabitoPublished: April 28, 2020 Explor Target Antitumor Ther. 2020;1:117–130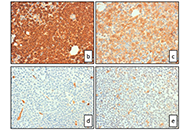 Multiple adverse drug reactions during all-trans retinoic acid treatment for acute promyelocytic leukemia: differentiation syndrome, bradycardia, intestinal necrosisOpen AccessCase ReportAll-trans retinoic acid (ATRA) induces complete remission in a high proportion of acute promyelocytic leukemia (APL). Nevertheless it is be associated with adverse drug reactions that might be life- [...] Read more.Valeria Ferla ... Nicola Stefano FracchiollaPublished: April 28, 2020 Explor Target Antitumor Ther. 2020;1:109–116
Multiple adverse drug reactions during all-trans retinoic acid treatment for acute promyelocytic leukemia: differentiation syndrome, bradycardia, intestinal necrosisOpen AccessCase ReportAll-trans retinoic acid (ATRA) induces complete remission in a high proportion of acute promyelocytic leukemia (APL). Nevertheless it is be associated with adverse drug reactions that might be life- [...] Read more.Valeria Ferla ... Nicola Stefano FracchiollaPublished: April 28, 2020 Explor Target Antitumor Ther. 2020;1:109–116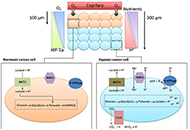 The impact of tumour pH on cancer progression: strategies for clinical interventionOpen AccessReviewDysregulation of cellular pH is frequent in solid tumours and provides potential opportunities for therapeutic intervention. The acidic microenvironment within a tumour can promote migration, invasi [...] Read more.Carol Ward ... Simon P LangdonPublished: April 28, 2020 Explor Target Antitumor Ther. 2020;1:71–100
The impact of tumour pH on cancer progression: strategies for clinical interventionOpen AccessReviewDysregulation of cellular pH is frequent in solid tumours and provides potential opportunities for therapeutic intervention. The acidic microenvironment within a tumour can promote migration, invasi [...] Read more.Carol Ward ... Simon P LangdonPublished: April 28, 2020 Explor Target Antitumor Ther. 2020;1:71–100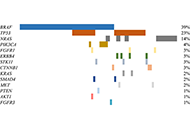 Implementation of an NGS panel for clinical practice in paraffin-embedded tissue samples from locally advanced and metastatic melanoma patientsOpen AccessOriginal ArticleAim: Single biomarker diagnostic test of BRAFV600 locus in metastatic melanoma is mandatory for treatment decision; however, multiple-gene based techniques, such as targeted next-generation sequenc [...] Read more.Paola Castillo ... Cristina TeixidoPublished: April 28, 2020 Explor Target Antitumor Ther. 2020;1:101–108
Implementation of an NGS panel for clinical practice in paraffin-embedded tissue samples from locally advanced and metastatic melanoma patientsOpen AccessOriginal ArticleAim: Single biomarker diagnostic test of BRAFV600 locus in metastatic melanoma is mandatory for treatment decision; however, multiple-gene based techniques, such as targeted next-generation sequenc [...] Read more.Paola Castillo ... Cristina TeixidoPublished: April 28, 2020 Explor Target Antitumor Ther. 2020;1:101–108